The genus trace: a function that shows values of genus (vertical axis) for subchains spanned between the first residue, and all other residues (shown on horizontal axis). The number of the latter residue and the genus of a given subchain are shown interactively.
Total Genus |
19
|
sequence length |
117
|
structure length |
117
|
Chain Sequence |
AGHMATTKRVLYVGGLAEEVDDKVLHAAFIPFGDITDIQIPLDYETEKHRGFAFVEFELAEDAAAAIDNMNESELFGRTIRVNLAKPMRIKEGSSRPVWSDDDWLKKFSGKTLEENK
|
The genus matrix. At position (x,y) a genus value for a subchain spanned between x’th and y’th residue is shown. Values of the genus are represented by color, according to the scale given on the right.
After clicking on a point (x,y) in the genus matrix above, a subchain from x to y is shown in color.
| publication title |
RNA binding induces an allosteric switch in Cyp33 to repress MLL1-mediated transcription.
pubmed doi rcsb |
| molecule tags |
Transcription
|
| source organism |
Homo sapiens
|
| molecule keywords |
Peptidyl-prolyl cis-trans isomerase E
|
| total genus |
19
|
| structure length |
117
|
| sequence length |
117
|
| ec nomenclature |
ec
5.2.1.8: peptidylprolyl isomerase. |
| pdb deposition date | 2022-03-31 |
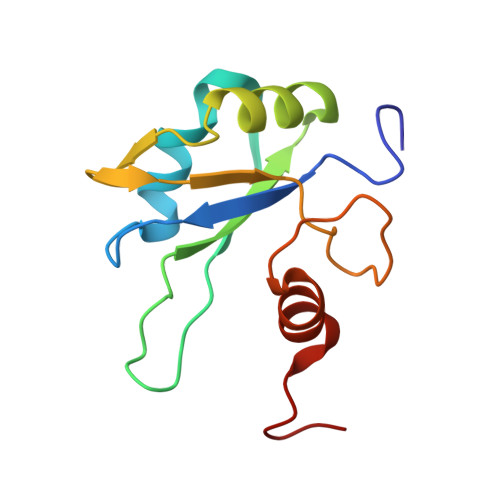
 Image from the rcsb pdb (www.rcsb.org)
Image from the rcsb pdb (www.rcsb.org)
#similar chains in the Genus database (?% sequence similarity) ...loading similar chains, please wait... #similar chains, but unknotted ...loading similar chains, please wait... #similar chains in the pdb database (?% sequence similarity) ...loading similar chains, please wait...